There's something new to be learnt everyday as a Biomedical Scientist. This is a record of my continous professional development .
About Me

- Clare
- I'm a busy Mum and a Biomedical Scientist in Haematology. My particular interest is in blood cell morphology and parasitology, where I never stop learning.
Monday, 20 November 2023
Which Parameters are truly useful in XbarM analysis and which are a waste of time!
 I'm a busy Mum and a Biomedical Scientist in Haematology.
My particular interest is in blood cell morphology and parasitology, where I never stop learning.
I'm a busy Mum and a Biomedical Scientist in Haematology.
My particular interest is in blood cell morphology and parasitology, where I never stop learning.
Tuesday, 26 September 2023
Unstable Haemoglobin Causing Inaccurate White Cell Differential on Sysmex XN
Look at these white cell differentials analysed on the
Sysmex XN. The first run to the right labelled run 1, gives a neutrophil count of
1.21 x 109/L and Eosinophils of 8.5 x 109/l. The white cell differential on second run
is completely reversed.
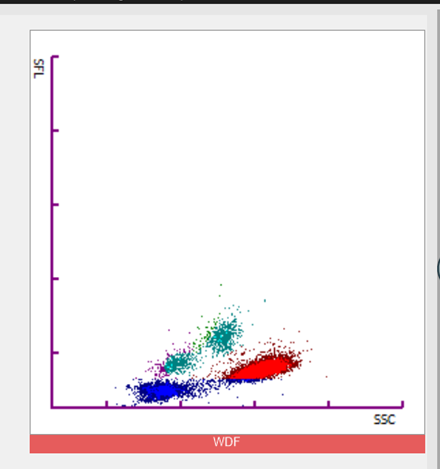
This can also be clearly seen on the WDF plots. On run 1, the large orange population of white cells were counted as eosinophils. On the second run, they were counted as
neutrophils represented by the blue population.
 |
| Run 1 |
 |
| Run 2 |
It is also clear that the white cell populations are
‘squashed’ and restricted to lower portion of the plot. Usually all populations would be
higher up the plot, with the monocytes being the highest . This is because side flouorescent
light (SFL) corresponds to the amount of DNA and RNA in a cell and monocytes have more than other normal white cells.
The inaccuracy of the white cell differential here, was caused by an unstable haemoglobin. These are haemoglobinopathies where the solubility of haemoglobin is altered, leading to precipitates of haemoglobin and haemolytic anaemia due to reduced red cell lifespan.
The reason the unstable haemoglobin leads to inaccuracies in the white cell differential, is because it does not act in the same way as it’s stable
counterparts when the red cells are lysed with Lysercell WDF on the Sysmex XNs.. This is an
important step in the measurement of white blood cells. Reading up on this White_Paper_RBC_Diseases.pdf
(sysmex-europe.com), it is suggested that the unstable haemoglobin
interferes with the fluorescence marker used to differentiate white blood cell
populations, leading to decreased flouorescence and inaccurate differentials. This is why the entire plot is shifted down and appears that all cells have little DNA or RNA content.
Therefore if you encounter an abnormal plot where the
fluorescence is reduced, it is firstly important to perform a manual
differential. Secondly there should be the suspicion of an unstable haemoglobin particularly if haemolysis is present and HPLC and or/genetic sequencing would be indicated.
 I'm a busy Mum and a Biomedical Scientist in Haematology.
My particular interest is in blood cell morphology and parasitology, where I never stop learning.
I'm a busy Mum and a Biomedical Scientist in Haematology.
My particular interest is in blood cell morphology and parasitology, where I never stop learning.
-
Zini et al, 2012 I recently reviewed the ICSH recommend...
-
I asked a colleague who is very experienced in thrombophilia testing and interpretation, how I could improve my…er.....not so good knowled...
-
Reviewing the XbarM analysis parameters used in our laboratory XbarM control program and looking at whether the current parameters being mon...





